| ||||
 Applying to the Schatzlab The Schatzlab is a highly interactive research group known for outstanding research in computational biology. We are developing the computational models and systems needed for understanding the causes of diseases, for developing better foods and biofuels, and for unlocking the secrets of the processes of life. This work depends on having a team of highly motivated postdocs, graduate students, undergraduate students, and bioinformatics engineers to explore all the different facets of these challenges. We will work together to establish and execute a premier research program in computational biology through the entire research life cycle: selecting research topics, establishing milestones, developing software and methods, analyzing data, writing papers & presenting your work. As you mature in the lab, you will be expected to take on more independence & responsibility so that you will be well prepared for your entire career path. Computational Post-Doctoral Researchers Applications are invited for a 2-3 year computational postdoctoral research position in the Schatz laboratory at Johns Hopkins University. The researcher will develop novel methods for large-scale DNA-seq, RNA-seq, and other genomics data related to human and/or plant genetics. Potential project include such as developing methods for discovering and characterizing somatic mutations related to cancer, or for assembling and analyzing large complex plant genomes. Ideal applicants will have:
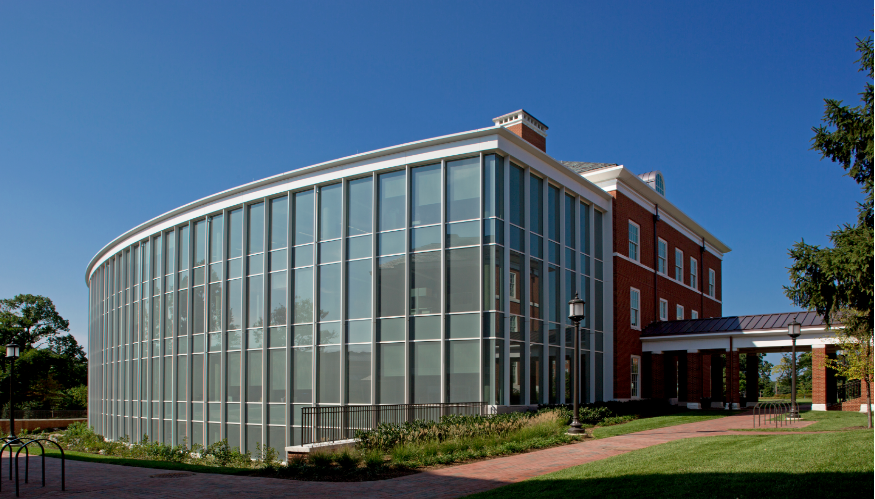
Graduate Students Graduate students interested in computational biology are encouraged to apply to either the Department of Computer Science or the Department of Biology at Johns Hopkins University. This pair of programs offers the greatest depth and breath on topics to establish a strong foundation for your thesis research. More information on the programs is available here. All graduate students must have strong programming and analytical skills, along with excellent communication abilities. Undergraduate Students Undergraduate students are encouraged to apply to the NSF-sponsored Undergraduate Research Program at Cold Spring Harbor Laboratory (CSHL), a ten week paid research experience in Bioinformatics, Genetics & Genomics, Cellular and Molecular Biology, Plant Biology, Neuroscience, and Cancer Biology. Approximately 25 students a year are selected from around the world to work side-by-side with CSHL Investigators and gain hands-on experience. In the past few years, my students have developed novel computational and mathematical methods for analyzing, plant, animal, and human genomes, several of which have lead to significant publications. Facilities The Schatzlab is housed in the top floor of Malone Building in the Computer Science Department on the Homewood campus of Johns Hopkins University. We work very closely with other research groups within Computer Science, as well as groups within Biology, Biostatistics, Biomolecular Engineering, Medicine and Oncology. Schatzlab members have access to considerable computational resources, including the 20,000-core cluster at the Maryland Advanced Research Computing Center. The Schatzlab has access to over 1M core hours / year and over 100 TB of shared disk space on the system as well as a dedicated high-memory server with 1TB of RAM, 144 cores, and 168TB of local disk. The computational development works side-by-side with our experimentalist collaborators to probe deeply into a variety of biological systems and diseases. We make extensive use of all major sequencing platforms including Illumina, PacBio, Oxford Nanopore, 10X Genomics, and others.
|